January 23, 2026
123 (4) e2406949123
Significance
The human topoisomerase IIIα–RMI1–RMI2 complex (TRR) is important for resolving entangled DNA structures known as ultrafine anaphase bridges (UFBs) during mitosis. Recent evidence suggests that this may involve the relaxation of negative supercoils generated by the protein PICH. Here, we report a single-molecule assay to measure supercoil relaxation in real time while imaging the binding of topoisomerases. Using this approach, we demonstrate that TRR can relax negative supercoils faster than they are expected to be generated by PICH. This deepens our mechanistic understanding of how TRR may facilitate UFB resolution. Moreover, we propose that our assay could be implemented to correlate protein dynamics with supercoil relaxation for other topoisomerase enzymes.
Abstract
Topoisomerase enzymes are essential for the regulation of DNA topology. Human topoisomerase IIIα is a Type 1A topoisomerase that exists as a complex with RMI1 and RMI2, known as TRR. The TRR complex can unlink entwined DNA strands and is known to be important for resolving DNA replication and recombination intermediates. It has recently been proposed that TRR can also relax transient negatively supercoiled loops of DNA generated by the translocase PICH and that this activity may help to facilitate the resolution of ultrafine anaphase bridges (UFBs) between segregating sister chromatids. However, the mechanism by which TRR interacts with, and processes, negatively supercoiled DNA is not well understood. Here, we establish a single-molecule strategy to simultaneously measure real-time changes in supercoiling density and visualize the interactions of TRR with underwound DNA using a combination of optical tweezers and fluorescence imaging. We demonstrate that TRR relaxes highly negatively supercoiled DNA in a processive manner and that the timescale for relaxation is less than the expected lifetime of the negatively supercoiled loops generated by PICH. We also show that in the absence of free protein in solution, TRR remains bound to the DNA for long time periods after the torsional stress has been released. Our findings provide a mechanistic basis for how TRR can relax negative supercoils, consistent with its proposed role in UFB resolution. Moreover, our assay could also be widely applied to study the interactions of other families of topoisomerases with negatively supercoiled DNA.
Topoisomerase enzymes are essential for regulating the topological state of DNA during processes such as DNA replication, transcription, and recombination (1–3). They achieve this by creating a transient break in DNA, through which another segment of DNA can pass. Topoisomerases can be classified into two families, referred to as Type 1 and Type 2, depending on whether they cleave one strand or both strands of the DNA (4). Both families can be further divided into different subfamilies. Type 1 topoisomerases consist of three subfamilies, known as Type 1A, Type 1B, and Type 1C (4, 5). Type 1A subfamily members are found in all domains of life and often play key roles in either maintaining supercoiling homeostasis or untangling (i.e., decatenating) entwined DNA structures (6–9). These enzymes require the presence of single-stranded (ss)DNA for catalytic activity (10–12), which often arises in negatively supercoiled (i.e., underwound) DNA (13–17) as well as in several replication/recombination intermediates, such as hemicatenanes (18–22).
Type 1A topoisomerases exhibit a highly conserved toroidal structure and alter the topological state of DNA using an enzyme-bridged strand-passage mechanism (3, 7–9), involving the following steps. First, the enzyme cleaves the ssDNA backbone, creating a covalent bond with the 5′-end of the DNA and a noncovalent interaction with the 3′-end (23–26). This generates an enzyme-bridged gate in the ssDNA that can open and close due to conformational changes in the enzyme (27–32). Upon opening of the gate, a second DNA segment (referred to as the transfer or T-strand) is able to enter the central cavity of the toroidal fold of the enzyme (28, 32). Once the second strand enters the cavity, the gate closes and the ssDNA backbone is religated, resulting in a change in the DNA linking number (Lk) of one (33–35).
Apart from reverse gyrases (36), Type 1A topoisomerases are ATP-independent and are often classified into two main subgroups, TopoI and TopoIII (4), named after the Type 1A topoisomerases in Escherichia coli. TopoI enzymes are, in general, more efficient at relaxing supercoils than (de)catenating DNA, whereas the opposite is typically the case for TopoIII enzymes (11, 37, 38). While many bacteria possess both TopoI and TopoIII, eukaryotes have only TopoIII (4), which often exists in complex with one or more OB-fold regulatory protein(s) (39–42). Higher eukaryotes typically encode two TopoIII enzymes, known as TopoIIIα and TopoIIIβ (4). TopoIIIα forms a stable complex with the OB-fold proteins RMI1 and RMI2, known as TRR (40, 41, 43). TRR can (de)catenate DNA efficiently and, together with the helicase BLM, it plays an important role in the dissolution of double Holliday junctions (19, 44–48). In addition, the TRR complex has been proposed to help resolve precatenane structures generated during replication (32).
TRR is also important for resolving ultrafine anaphase bridges (UFBs), which are entangled DNA structures that bridge the separating daughter chromosomes during anaphase (49, 50). The mechanism by which TRR contributes to UFB resolution is thought to depend on the type of UFB that is present (7, 21). For example, UFBs arising from common fragile sites likely contain incompletely replicated DNA, consisting of both double-stranded (ds)DNA and ssDNA (21). It has been proposed that TRR (acting in concert with BLM) plays a direct role in resolving these UFBs due to its abilities to bind to ssDNA and to decatenate DNA (21, 32). In contrast, UFBs linked to centromeres typically consist of fully catenated dsDNA (21). Ultimately, it is believed that the Type 2 topoisomerase TopoIIα is responsible for decatenating centromeric UFBs (21, 51–53). However, there is recent evidence that TRR may also contribute to this pathway, by cooperating with the protein PICH (7, 54), which is an ATP-dependent dsDNA translocase that is found to localize to UFBs (49, 50, 55). Recent studies have shown that the translocation activity of PICH can result in the extrusion of transient dsDNA loops (54, 56). In a torsionally constrained substrate, these loops consist of (hyper)negatively supercoiled DNA, and thus, the DNA adjacent to the loops is positively supercoiled (54). It has been demonstrated that Type 1A topoisomerases (including TRR) can relax the negative supercoils in the loops generated by PICH. This concerted action results in a substrate that is, overall, positively supercoiled. Since positively supercoiled DNA is the preferred substrate for TopoIIα (57, 58), it has been hypothesized that TRR-mediated relaxation of the negatively supercoiled loops generated by PICH may play an important role in the resolution of centromeric UFBs (7, 54).
A recent study using magnetic tweezers has demonstrated that the catalytic cycle of human TopoIIIα exhibits long pause times, during which the enzyme–ssDNA complex is thought to exist in an open, cleaved state (59). Further, it was shown that RMI1 enhances the rate of TopoIIIα binding to ssDNA and acts to stabilize the open, cleaved complex, resulting in longer pause times (59). Nonetheless, the mechanisms by which TRR interacts with and processes negatively supercoiled DNA are still not well understood, especially in the context of its proposed role in UFB resolution.
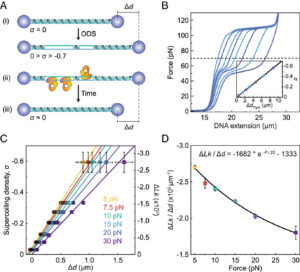
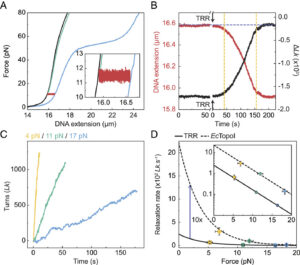
Here, we report a single-molecule assay to probe the interaction of topoisomerases with negatively supercoiled DNA, which we apply to study the mechanism of supercoil relaxation by human TRR. Our assay exploits a recently developed approach called Optical DNA Supercoiling (ODS), which enables rapid generation of negatively supercoiled DNA by using dual-trap optical tweezers (17, 60). ODS offers several advantages for the study of topoisomerase–DNA interactions. First, the negatively supercoiled substrate can be moved freely between different channels of a microfluidic flow cell, allowing rapid exchange of buffer/protein solutions. Second, ODS is readily compatible with fluorescence microscopy, offering the potential to directly visualize the binding of topoisomerases to the supercoiled DNA substrate. By applying our ODS-based assay, we demonstrate that TRR can relax hypernegatively supercoiled DNA in a highly processive manner, with single complexes capable of performing up to several thousand strand-passage reactions. Our study also indicates that the timescale for TRR-induced relaxation of hypernegatively supercoiled DNA is shorter than the previously reported average lifetime of negatively supercoiled loops generated by PICH (54). These findings provide a rationale for how TRR may relax negatively supercoiled DNA in the context of UFBs.
Link to the full PNAS article here